Description
The DeepChek®Assay – HCV NS3 Drug Resistance V1 (RUO) is intended to be used for HCV drug resistance assessment. It provides drug susceptibility information for viral NS3 inhibitors. It combines target-specific PCR reagents with in vitro diagnostic software both compatible with either Sanger or Next Generation Sequencing platforms.
Assay should be used for patients with documented HCV genotype 1 to 6 (pan-genotypic assay).
Methodology
DNA Sequencing • Reverse Transcriptase Polymerase Chain Reaction (RT-PCR)
More information on the DeepChek® Assays – Click here
More information on the DeepChek® Software – Click here
Characteristics and performances
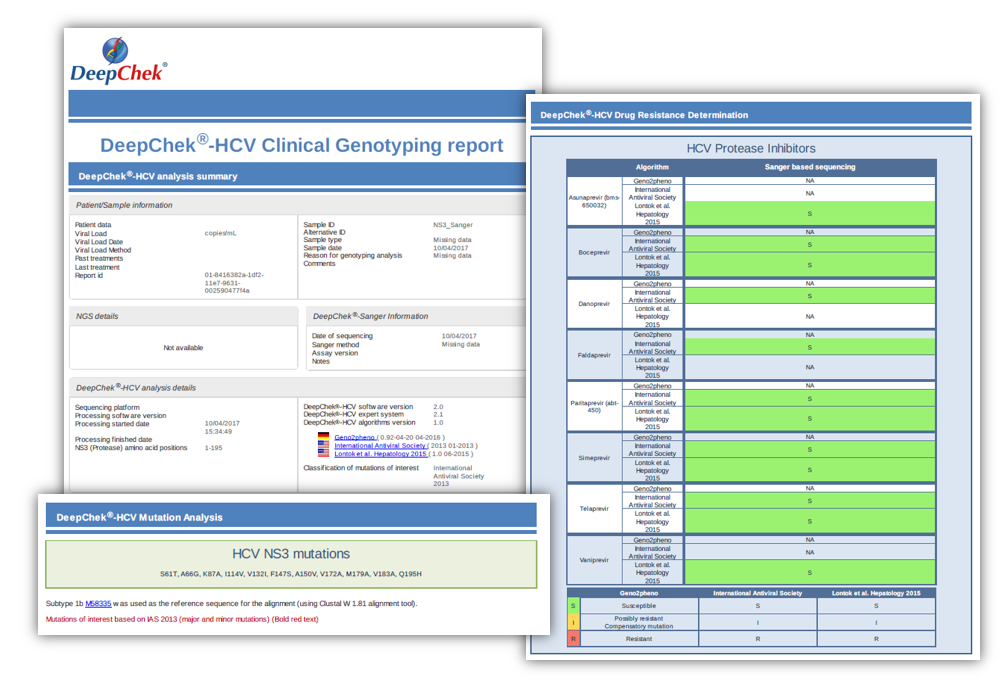
Examples of reports
For SANGER sequencing
For NGS sequencing
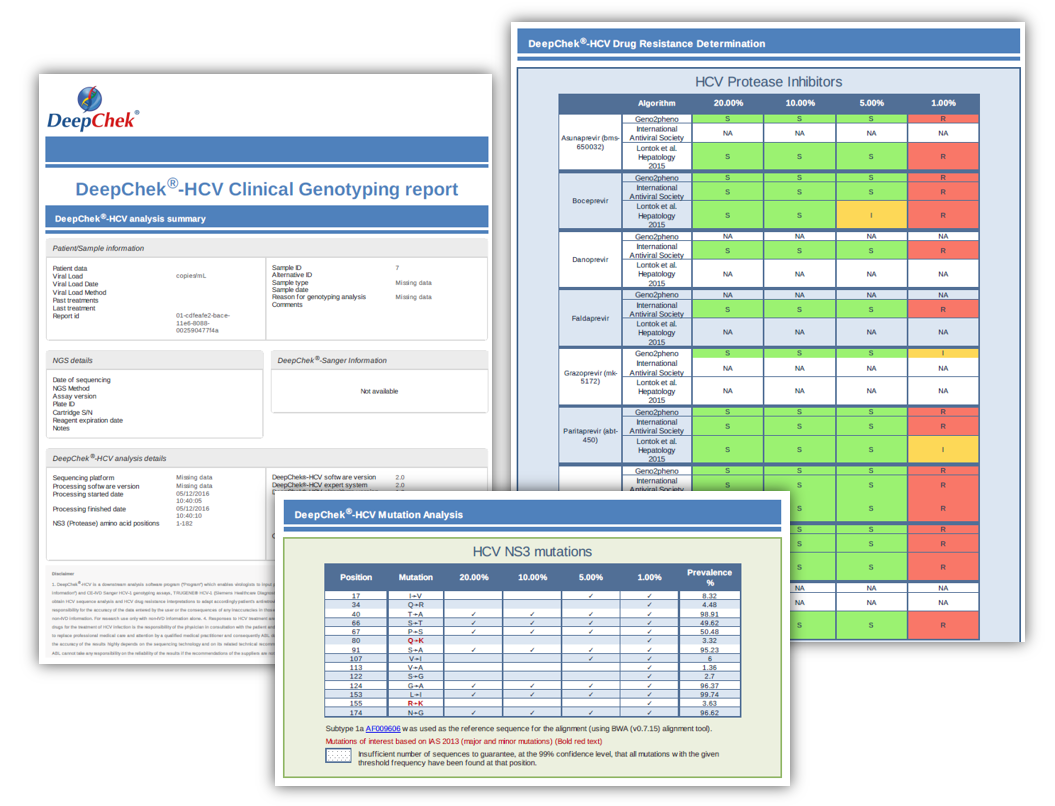

Ordering information
Downloads
General documentation
-
Protocol
-
Installation check list
-
Q&A
MSDS
-
International