Description
The DeepChek®-HCV Assay NS5B Drug Resistance V4 (RUO) Drug Resistance Assay is intended to be used for HCV drug resistance assessment. It provides drug susceptibility information for viral NS5B inhibitors. It combines target-specific PCR reagents with in vitro diagnostic software both compatible with either Sanger or Next Generation Sequencing platforms.
Assay should be used for patients with documented HCV genotype 1 to 6 (pan-genotypic assay).
Methodology
DNA Sequencing • Reverse Transcriptase Polymerase Chain Reaction (RT-PCR)
More information on the DeepChek® Assays – Click here
More information on the DeepChek® Software – Click here
Characteristics and performances
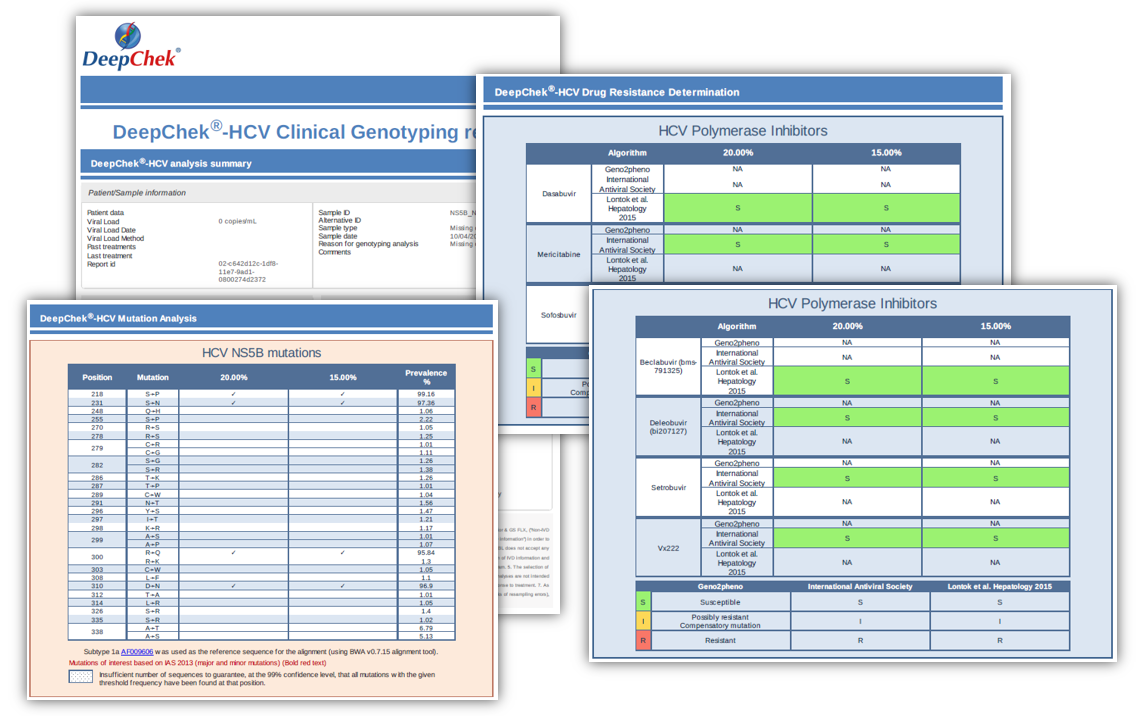
Examples of reports

Ordering information
Downloads
General documentation
-
Protocol
-
Installation check list
-
Q&A
MSDS
-
International