Description
The DeepChek® Assay-HCV NS5A Drug Resistance (RUO) is intended to be used for HCV drug resistance assessment. It provides drug susceptibility information for viral NS5A inhibitors. It combines target-specific PCR reagents with in vitro diagnostic software both compatible with either Sanger or Next Generation Sequencing platforms.
Assay should be used for patients with documented HCV genotype 1 to 6 (pan-genotypic assay).
For genotype 2 samples, in case of failure, please use the DeepChek® Assay NS5A (GT2) Drug Resistance V1 Assay(106A24) .
Methodology
DNA Sequencing • Reverse Transcriptase Polymerase Chain Reaction (RT-PCR)
Clinical Significance
EASL Recommendations on Treatment of Hepatitis C 2016: “Physicians who have easy access to a reliable test assessing HCV resistance to NS5A inhibitors (spanning amino acids 24 to 93) can use these results to guide their decisions, as specified in these recommendations. The test should be based on population sequencing (reporting RASs as “present” or “absent”) or deep sequencing with a cut-off of 15% (only RASs that are present in more than 15% of the sequences generated must be considered)”.
More information on the DeepChek® Assays – Click here
More information on the DeepChek® Software – Click here
Characteristics and performances
Examples of reports
For SANGER sequencing
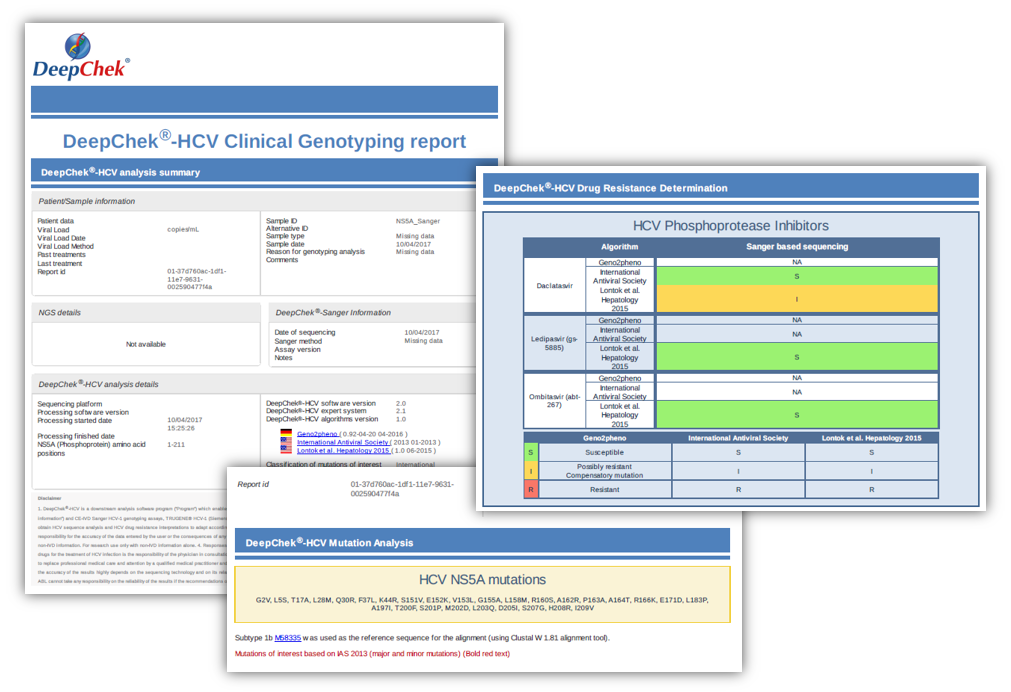
For NGS sequencing
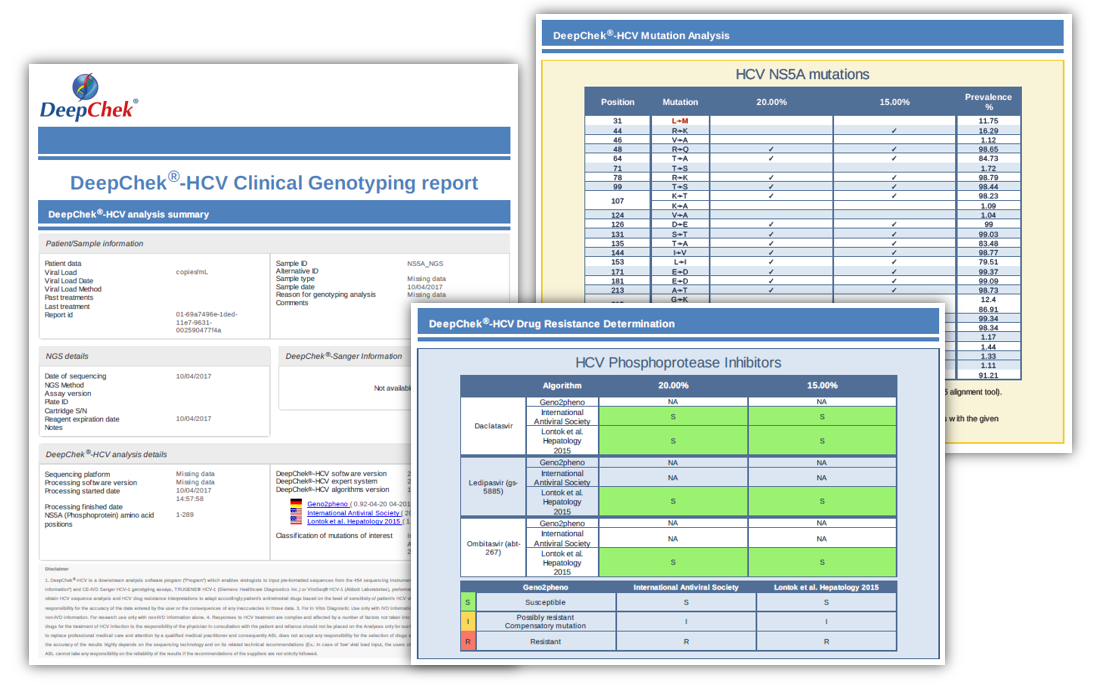

Ordering information
Downloads
General documentation
-
Protocol
-
Installation check list
-
Q&A
MSDS
-
International
Posters
-
Retreatment with Direct Active Antivirals of Genotype 1, 3 and 4 Chronic Hepatitis C Patients who Previously Failed an Anti-NS5A-Containing Regimen in Real World